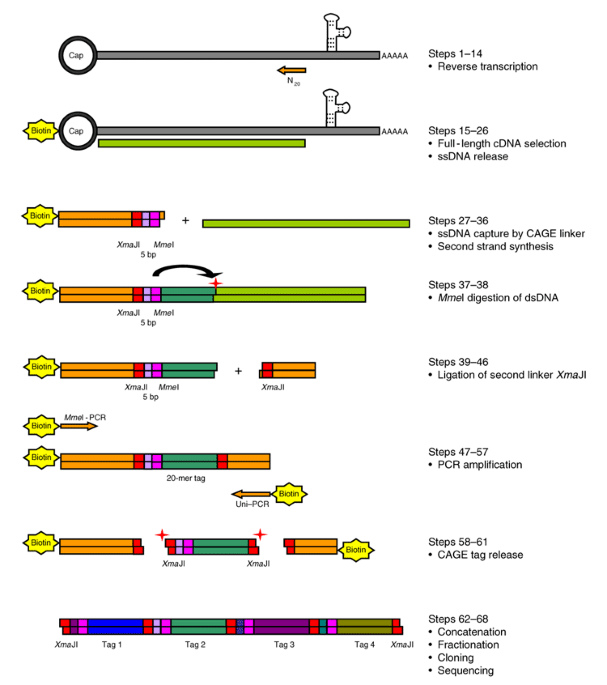
Cap analysis of gene expression (CAGE) and noncoding regulatory elements | Seminars in Immunopathology

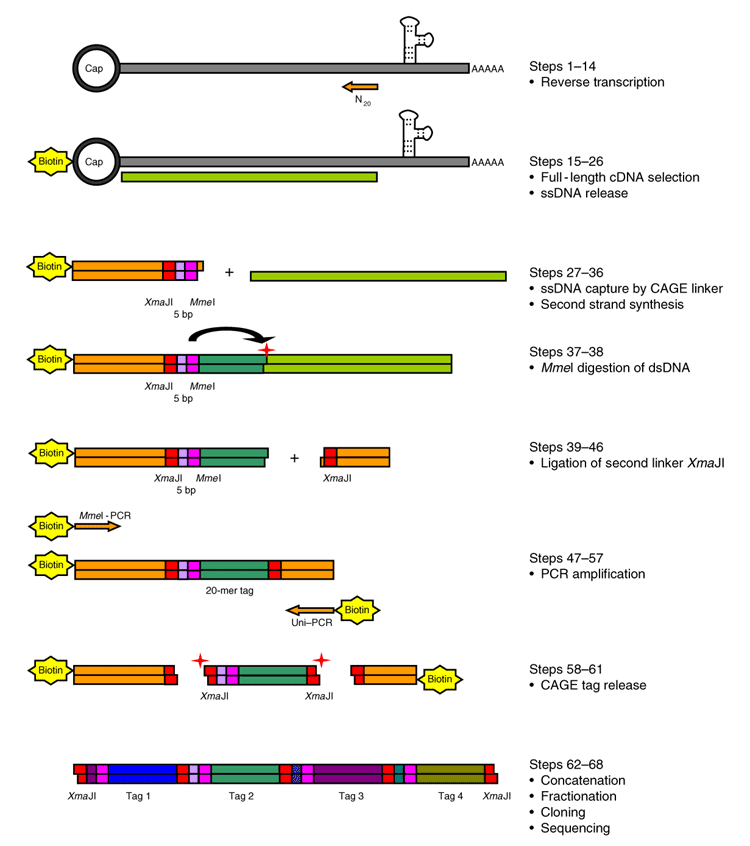
Cap analysis gene expression for high-throughput analysis of transcriptional starting point and identification of promoter usage | PNAS
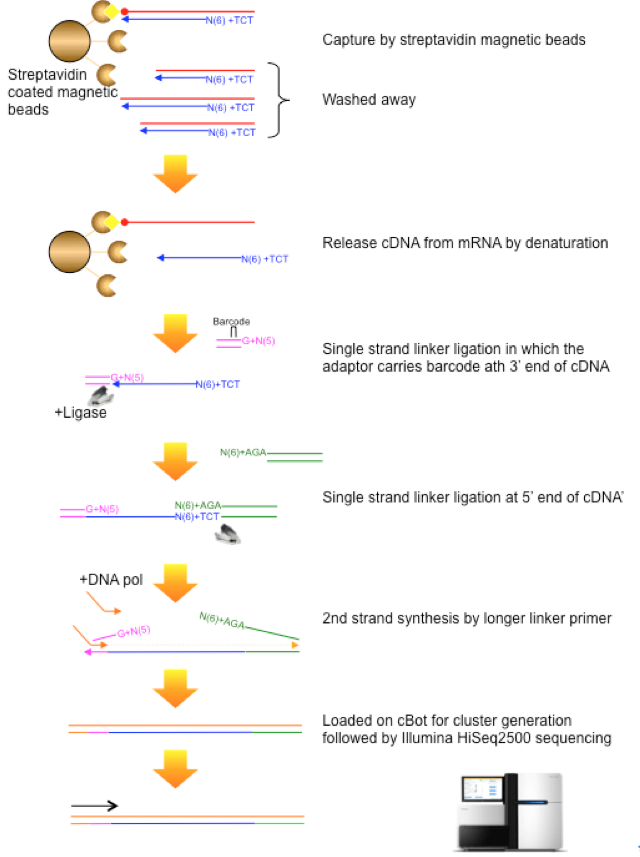
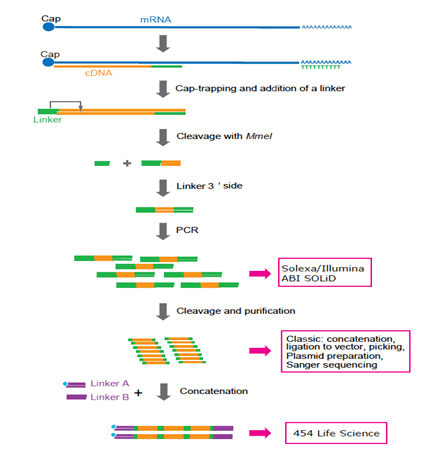
CAGE : A method for genome-wide identification of transcription start sites | 理化学研究所 ライフサイエンス技術基盤研究センター
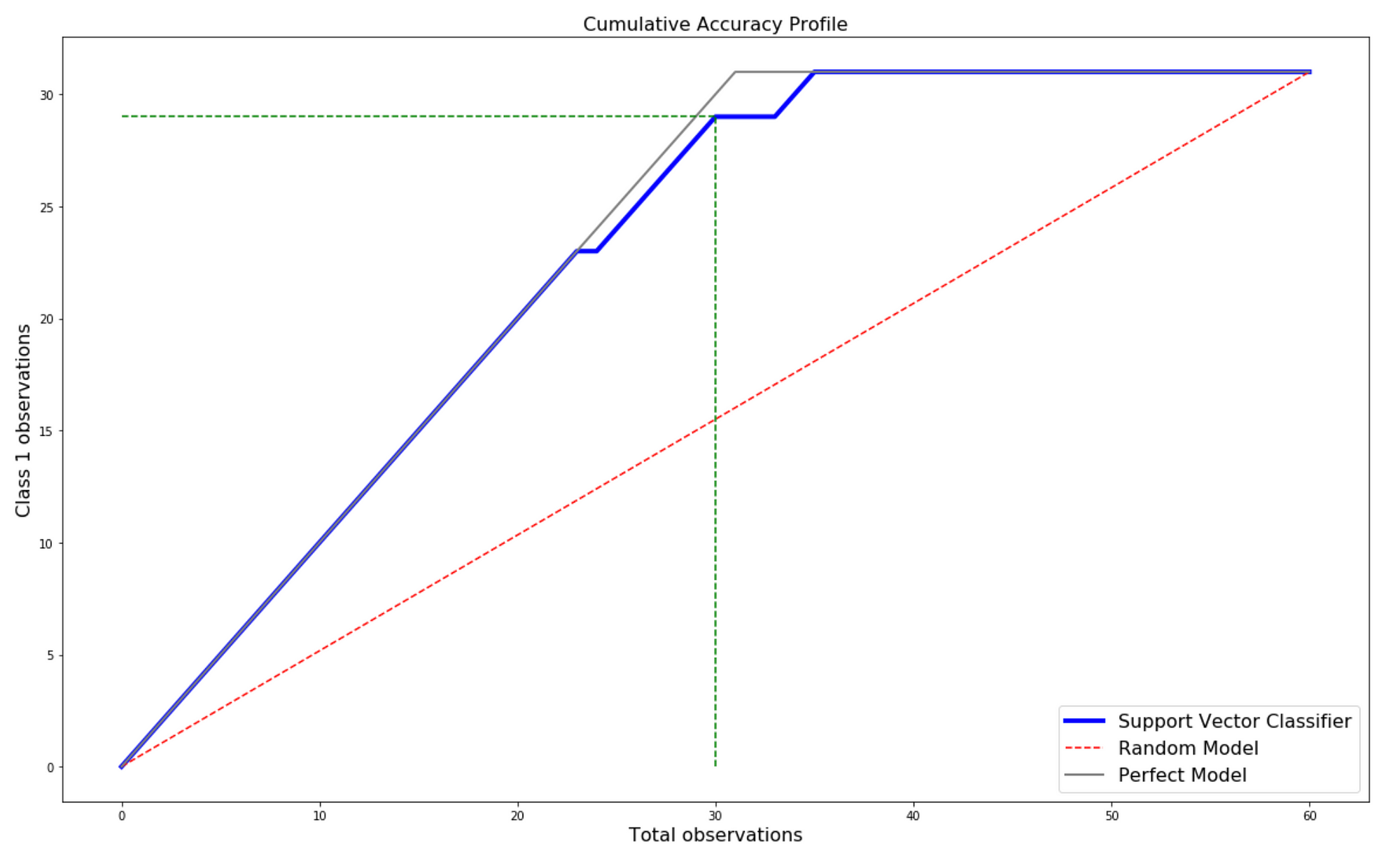
Machine Learning Classifier evaluation using ROC and CAP Curves | by Karan Bhanot | Towards Data Science

Canonical analysis of principal coordinates (CAP) ordination plot (Bray-Curtis) of fish assemblage data for each experimental treatment at each geographic location.
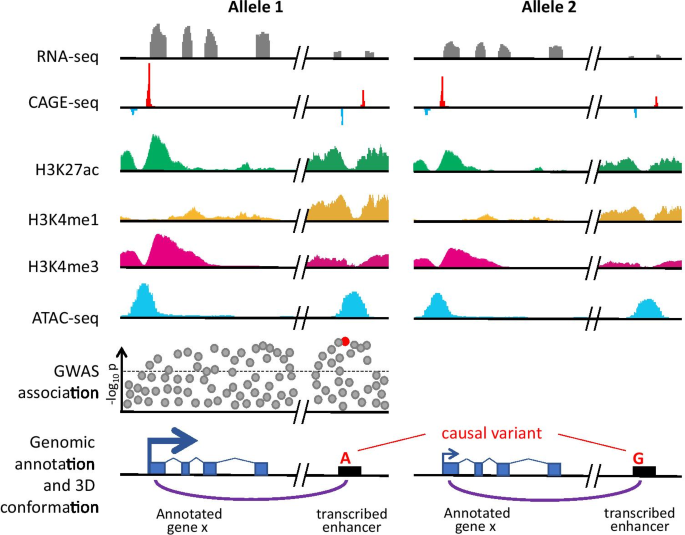
Cap analysis of gene expression (CAGE) and noncoding regulatory elements | Seminars in Immunopathology
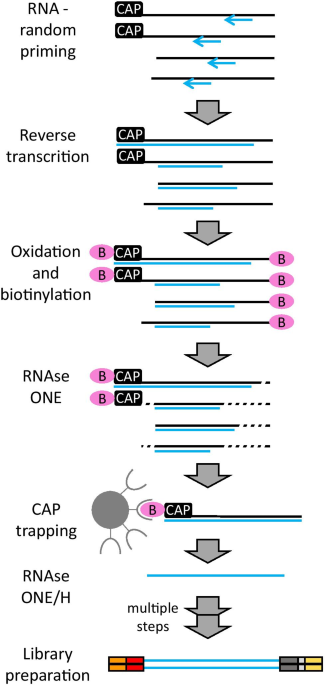
Cap analysis of gene expression (CAGE) and noncoding regulatory elements | Seminars in Immunopathology
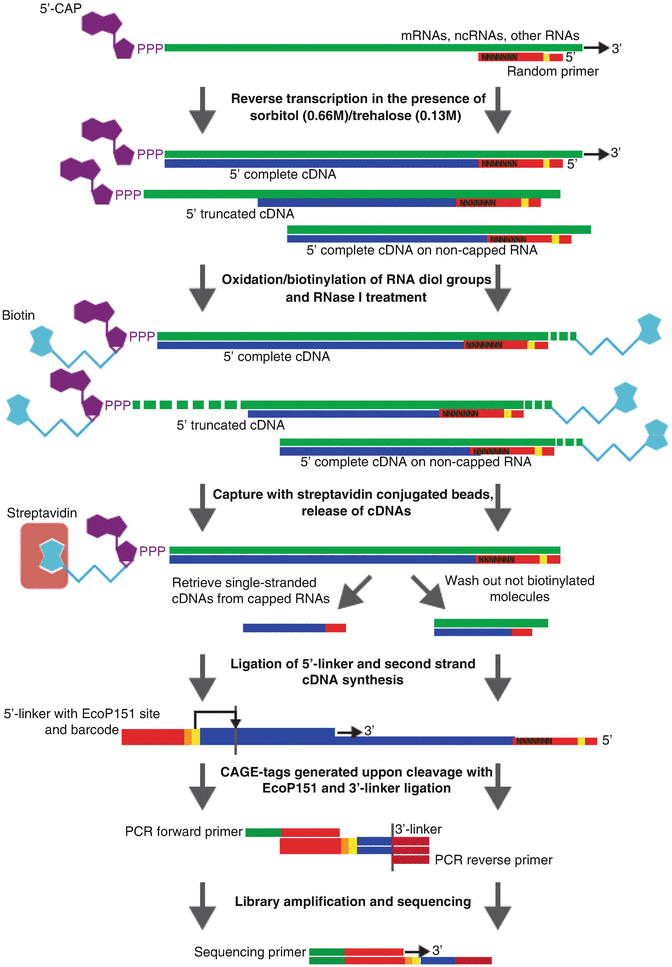
Deep Cap Analysis of Gene Expression (CAGE): Genome-Wide Identification of Promoters, Quantification of Their Activity, and Transcriptional Network Inference | SpringerLink
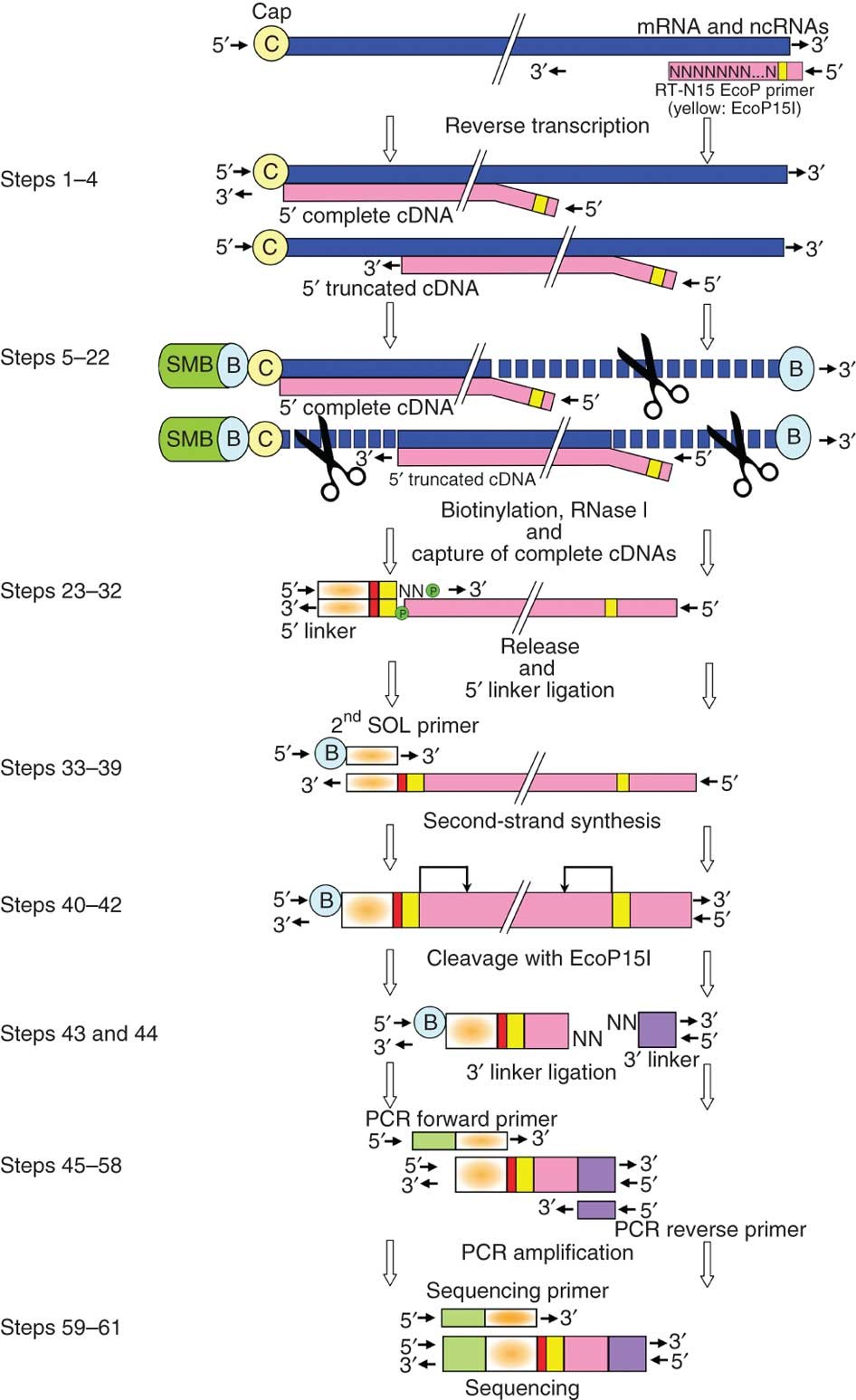
5′ end–centered expression profiling using cap-analysis gene expression and next-generation sequencing | Nature Protocols